# Optional: run once if you want to use Levene's test
# install.packages("car")F-Test and ANOVA
C5025HF
How to use this guide
This guide is written for you. You’ll read short explanations, then immediately run R code yourself to reproduce the results. Every step shows both the code and the output you should expect.
- By the end, you will be able to:
- Explain what ANOVA tests and when to use it.
- Fit a one-way ANOVA in R and interpret the output.
- Check model assumptions with plots and simple tests.
- Run and interpret post‑hoc comparisons (Tukey HSD and adjusted t‑tests).
- Understand the idea of two‑way ANOVA (factor + blocks, interaction).
Setup
You only need base R for everything in this guide. One optional test uses the car package.
If you use RStudio: open a new R script or Quarto document, and copy/paste the code chunks as you go.
Origins of the F‑test
- William Sealy Gosset (pen name “Student”) developed the small‑sample t‑test while working at Guinness to improve quality control.
- Ronald A. Fisher formalised variance partitioning and introduced the F‑statistic for comparing mean square quantities.
- The F‑distribution is associated with Fisher and popularised by Snedecor (hence the letter “F”).
- Big idea: compare explained variation to unexplained variation using a ratio of mean squares.
From t to F: why a ratio?
- If a model explains real structure, the “between” mean square should exceed the “within” mean square.
- Under a true null, both mean squares estimate the same variance → their ratio should be near 1.
- With normal errors, these scaled sums of squares follow chi‑squared distributions; their ratio follows an F‑distribution with appropriate degrees of freedom.
Where the F‑test appears
- One‑way and multi‑factor ANOVA global tests (“are any means different?”).
- Regression: overall model significance test.
- Standalone variance comparison: test whether two normal populations have equal variances.
Standalone F‑test example: comparing two variances
We’ll simulate two normal samples with different variances and test \(\sigma_x^2 = \sigma_y^2\).
set.seed(42)
x <- rnorm(30, mean = 0, sd = 1.0)
y <- rnorm(30, mean = 0, sd = 1.6)
var.test(x, y) # F-test for equality of variances
F test to compare two variances
data: x and y
F = 0.55801, num df = 29, denom df = 29, p-value = 0.122
alternative hypothesis: true ratio of variances is not equal to 1
95 percent confidence interval:
0.2655929 1.1723758
sample estimates:
ratio of variances
0.5580096 How to interpret:
H0: the population variances are equal. A small p‑value suggests different variances.
This test is sensitive to non‑normality; use with caution if distributions are heavy‑tailed or skewed.
What problem does ANOVA solve?
When you compare more than two group means (e.g., yields under fertilisers A, B, C), running many t‑tests inflates the chance of a false positive (Type I error). ANOVA answers one global question:
- \(H_0\): \(\mu_1 = \mu_2 = \dots = \mu_k\) (all group means equal)
- \(H_1\): at least one mean differs
It uses a single F‑statistic to compare “between‑groups” variation to “within‑groups” variation.
From variation to sums of squares
- Total variation is partitioned into:
- Between‑groups (explained by the factor/treatment)
- Within‑groups (unexplained residual/error)
- Notation:
- \(SS_T\) = total sum of squares
- \(SS_B\) = between‑groups sum of squares (treatment)
- \(SS_W\) = within‑groups sum of squares (residual/error)
We convert sums of squares to mean squares by dividing by degrees of freedom:
- \(MS_B = SS_B / (k - 1)\), where (k) is the number of groups
- \(MS_W = SS_W / (N - k)\), where (N) is the total number of observations
The F‑statistic is the ratio: \(F = \frac{\text{MS}_B}{\text{MS}_W}\). If all group means are truly equal, the ratio tends to be near 1; larger values suggest group means differ.
Hands‑on: One‑way ANOVA in R
We’ll create a small dataset with one numeric response (yield) and one factor (fert).
Take a quick look:
yield fert
1 20 A
2 22 A
3 19 A
4 21 A
5 34 B
6 35 B yield fert
Min. :19.00 A:4
1st Qu.:21.75 B:4
Median :28.00 C:4
Mean :27.75
3rd Qu.:34.00
Max. :36.00
A B C
4 4 4 Optional: visualise group differences before modelling.
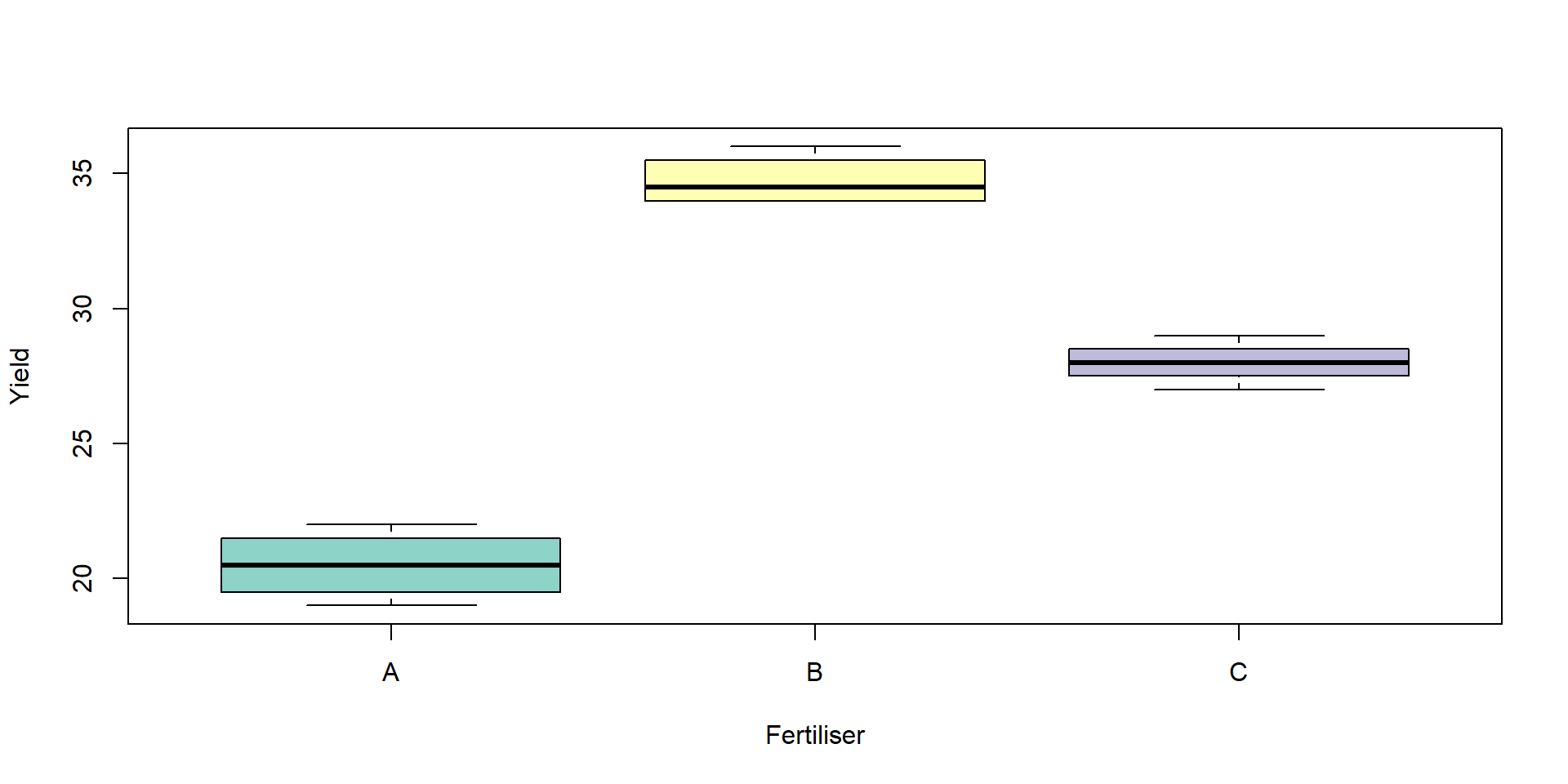
Fit the ANOVA model and view the ANOVA table:
Df Sum Sq Mean Sq F value Pr(>F)
fert 2 406.5 203.25 187.6 4.61e-08 ***
Residuals 9 9.7 1.08
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1How to read the table:
Df: degrees of freedom (treatment and residual)
Sum Sq: \(SS_B\) and \(SS_W\)
Mean Sq: \(MS_B\) and \(MS_W\)
F value: the test statistic \(F = \frac{\text{MS}_B}{\text{MS}_W}\)
Pr(>F): the p‑value (small p suggests at least one mean differs)
Manual ANOVA (by hand) and verification
Let’s compute the ANOVA components directly and confirm they match aov():
# Group means and counts
group_means <- tapply(fert$yield, fert$fert, mean)
group_counts <- table(fert$fert)
grand_mean <- mean(fert$yield)
# Between-groups sum of squares (SSB)
SSB <- sum(group_counts * (group_means - grand_mean)^2)
# Within-groups sum of squares (SSW)
SSW <- sum(unlist(tapply(fert$yield, fert$fert, function(v) sum((v - mean(v))^2))))
# Degrees of freedom
k <- length(group_means)
N <- nrow(fert)
df_between <- k - 1
df_within <- N - k
# Mean squares and F
MSB <- SSB / df_between
MSW <- SSW / df_within
F_manual <- MSB / MSW
# Assemble a manual ANOVA table
anova_manual <- data.frame(
Source = c("fert", "Residuals"),
Df = c(df_between, df_within),
`Sum Sq`= c(SSB, SSW),
`Mean Sq`= c(MSB, MSW),
check.names = FALSE
)
anova_manual Source Df Sum Sq Mean Sq
1 fert 2 406.50 203.250000
2 Residuals 9 9.75 1.083333[1] 187.6154Analysis of Variance Table
Response: yield
Df Sum Sq Mean Sq F value Pr(>F)
fert 2 406.50 203.250 187.62 4.607e-08 ***
Residuals 9 9.75 1.083
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1You should see identical sums of squares, degrees of freedom, mean squares, and F.
Assumptions you should check
ANOVA relies on:
Independence of observations (this is a design issue).
Approximately normal residuals.
Homogeneity of variances across groups.
The standard diagnostic plots are a good start:
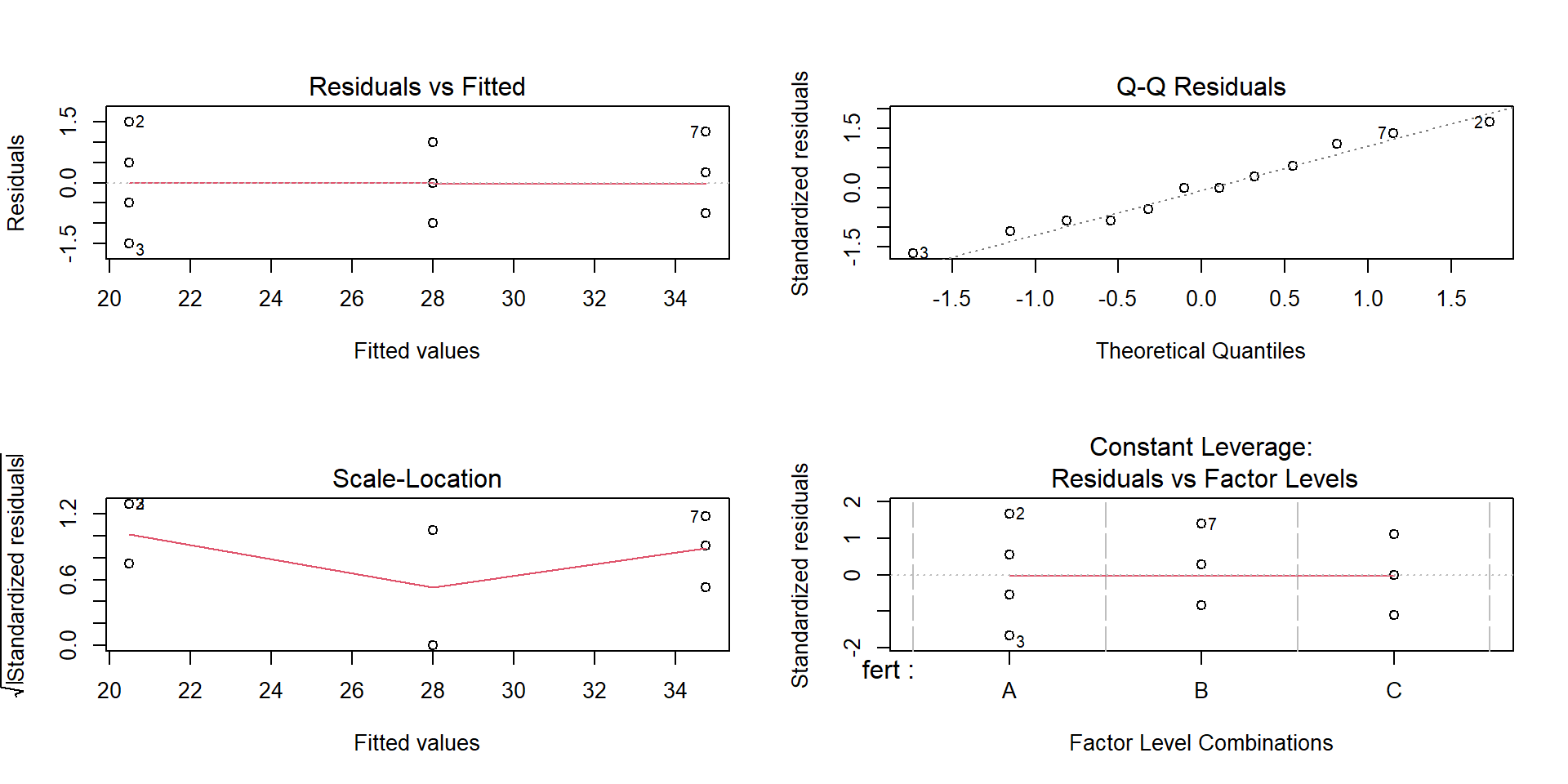
What to look for:
Residuals vs Fitted: no strong patterns or funnels.
Normal Q‑Q: points near a straight line.
Formal checks (optional and context‑dependent):
Shapiro-Wilk normality test
data: residuals(fit)
W = 0.96598, p-value = 0.8645# Homogeneity of variances (choose one)
bartlett.test(yield ~ fert, data = fert) # Bartlett's test (sensitive to non‑normality)
Bartlett test of homogeneity of variances
data: yield by fert
Bartlett's K-squared = 0.57949, df = 2, p-value = 0.7485If assumptions look poor: consider transforming the response (e.g., log), using a robust method, or revisiting the design/measurement process.
Interpreting the Q‑Q plot (in detail)
First, plot the residuals explicitly so you can inspect their shape.
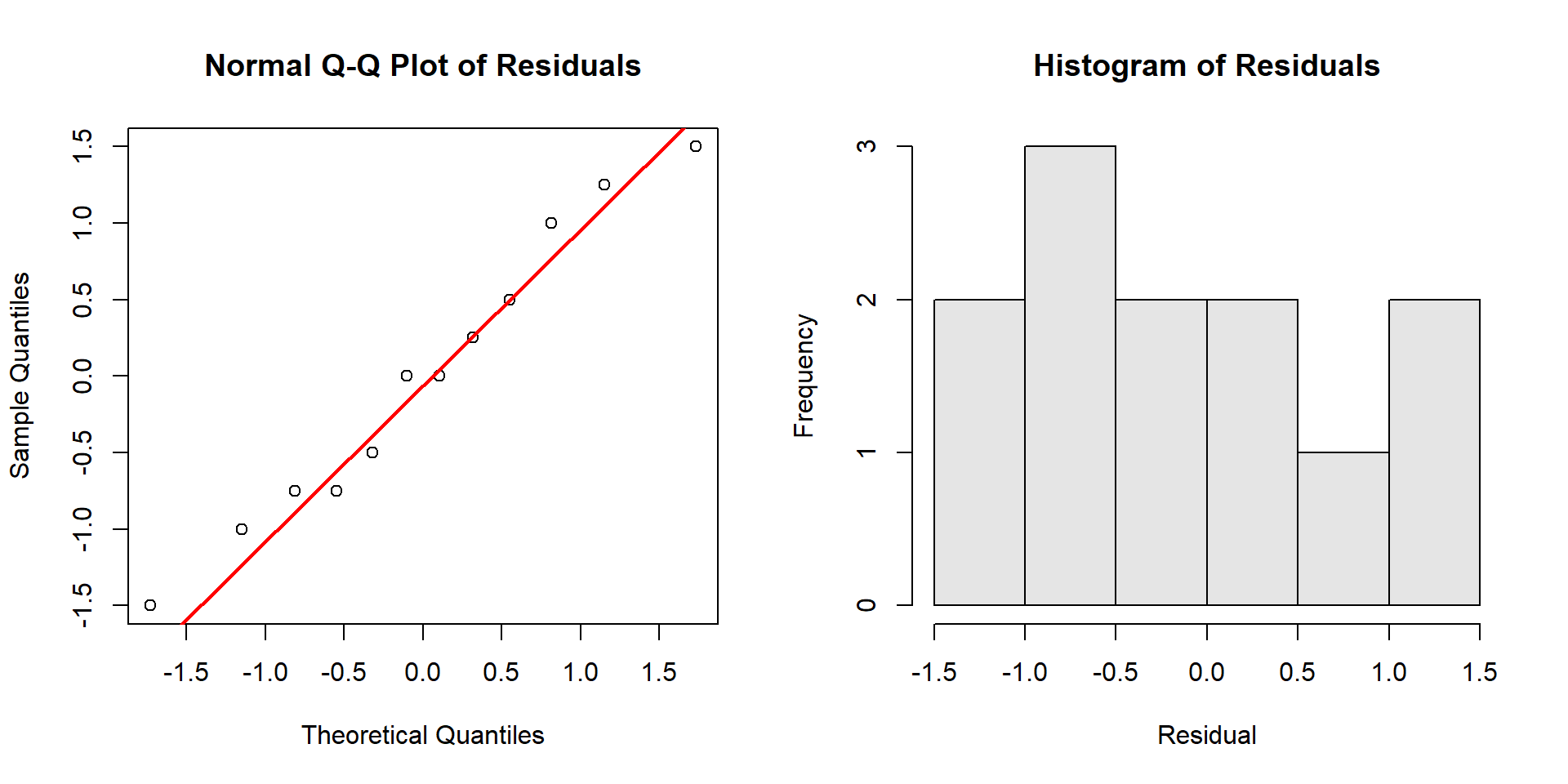
How to read the Q‑Q plot:
- Points close to the straight line → residuals are approximately normal.
- S‑shaped curve (points above the line on the left and below on the right, or vice versa) → skewness.
- Points that bow away at both ends → heavy tails (more extreme residuals than normal).
- Points that stay close in the middle but diverge strongly at ends → tail problems that can affect small‑sample inference.
What to do:
Minor deviations are usually fine; focus on large, systematic patterns.
Consider transformations (e.g., log) for right‑skewed positive data, or a different model if deviations reflect design issues.
Which means differ? Post‑hoc comparisons
ANOVA tells you there is a difference somewhere. To find where, use multiple comparisons that control familywise error.
Tukey’s Honest Significant Difference (HSD):
Tukey multiple comparisons of means
95% family-wise confidence level
Fit: aov(formula = yield ~ fert, data = fert)
$fert
diff lwr upr p adj
B-A 14.25 12.19514 16.30486 0.00e+00
C-A 7.50 5.44514 9.55486 8.20e-06
C-B -6.75 -8.80486 -4.69514 1.95e-05You can also visualise the Tukey intervals:
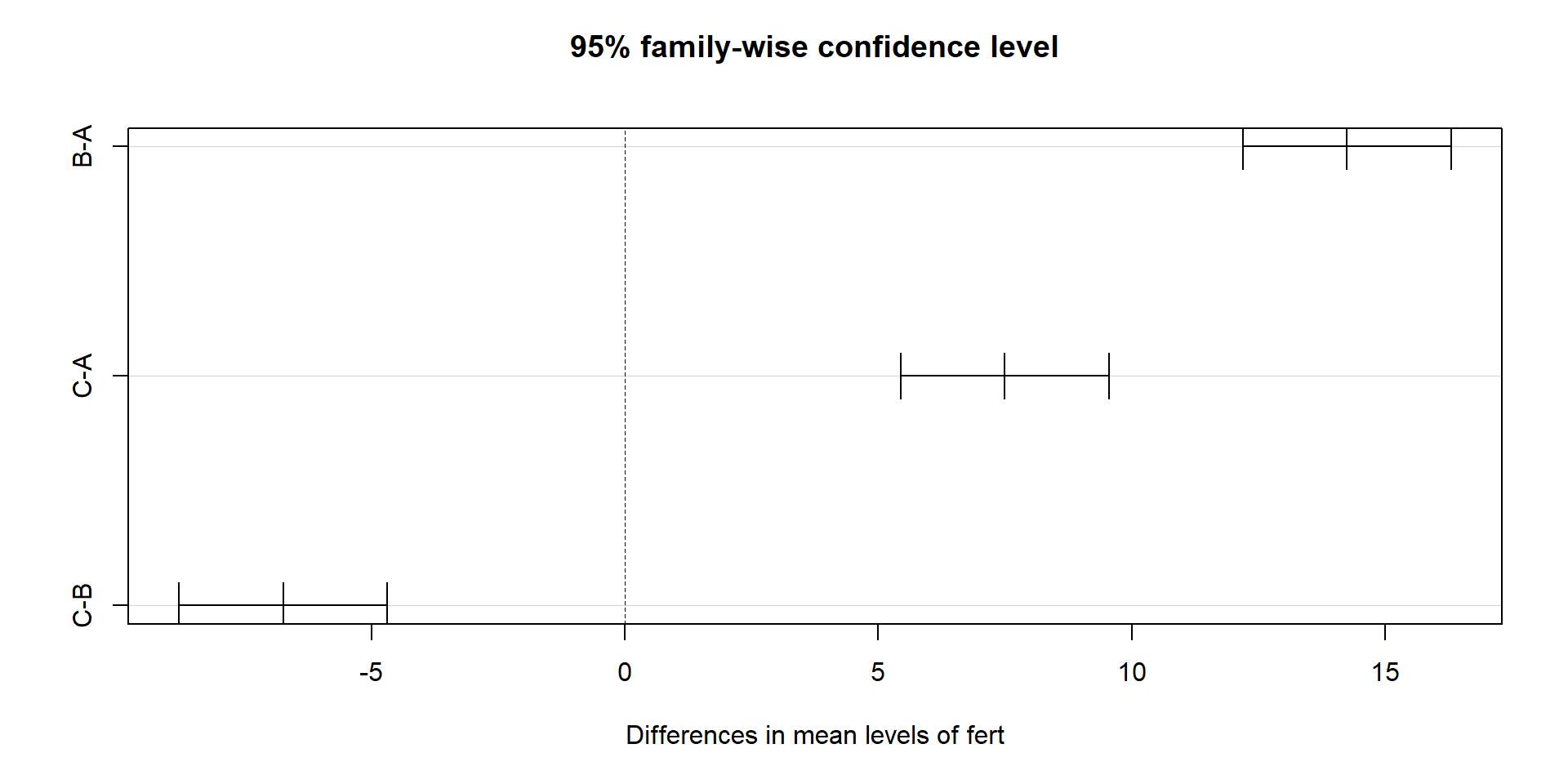
Pairwise t‑tests with p‑value adjustment (alternative approach):
Pairwise comparisons using t tests with pooled SD
data: fert$yield and fert$fert
A B
B 3.6e-08 -
C 9.2e-06 2.2e-05
P value adjustment method: bonferroni How to interpret:
Each pair shows a difference in means, a confidence interval (Tukey), and an adjusted p‑value.
If a confidence interval does not include 0, or the adjusted p‑value is small, that pair differs.
Reporting your result
Use clear, reproducible language. Replace the placeholders with your numbers:
“Yield differed by fertiliser treatment (one‑way ANOVA, F(df1, df2) = Fvalue, p = pvalue). Tukey’s HSD indicated that [list groups] differed.”
Example structure: - State the ANOVA result (F, degrees of freedom, p‑value). - Briefly summarise which pairs differ based on Tukey (with direction).
Extension: Two‑way ANOVA (factor + blocks)
Now you’ll add a blocking factor (e.g., position) to reduce unexplained variation.
set.seed(1)
twoway <- expand.grid(
fert = factor(c("A","B","C")),
block = factor(1:4)
)
twoway$yield <- c(20, 21, 19, 22,
34, 33, 36, 35,
27, 28, 30, 29) + rnorm(12, 0, 0.6)
tw_fit <- aov(yield ~ fert + block, data = twoway)
summary(tw_fit) Df Sum Sq Mean Sq F value Pr(>F)
fert 2 35.24 17.62 1.11 0.3889
block 3 296.19 98.73 6.22 0.0285 *
Residuals 6 95.23 15.87
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Interpretation:
fert: main treatment effect.block: accounts for position/batch effects.Residual: remaining noise.
You can also test for interaction (does the fertiliser effect depend on block?):
ANCOVA: ANOVA with a covariate
Why you need it:
Sometimes groups differ partly because of a continuous baseline variable (e.g., initial size, starting weight, pre‑test score).
ANCOVA adjusts group comparisons for that covariate, improving precision and providing “fair” comparisons at the same covariate level.
Conceptually, you fit parallel regression lines with a common slope across groups and compare their intercepts (adjusted means).
set.seed(2)
n_per <- 12
ancova <- data.frame(
fert = factor(rep(c("A","B","C"), each = n_per)),
size0 = rnorm(3 * n_per, mean = 10, sd = 2)
)
# True treatment effect and slope
beta0 <- 5; slope <- 0.8
trt_effect <- c(A = 0, B = 1.2, C = 2.0)
ancova$yield <- with(ancova, beta0 + trt_effect[fert] + slope * size0 + rnorm(nrow(ancova), 0, 1))
# Parallel slopes model (no interaction)
fit_ancova <- aov(yield ~ fert + size0, data = ancova)
summary(fit_ancova) Df Sum Sq Mean Sq F value Pr(>F)
fert 2 17.91 8.95 6.299 0.00494 **
size0 1 76.56 76.56 53.858 2.41e-08 ***
Residuals 32 45.49 1.42
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Optional: test for non-parallel slopes
fit_ancova_int <- aov(yield ~ fert * size0, data = ancova)
anova(fit_ancova, fit_ancova_int)Analysis of Variance Table
Model 1: yield ~ fert + size0
Model 2: yield ~ fert * size0
Res.Df RSS Df Sum of Sq F Pr(>F)
1 32 45.489
2 30 42.782 2 2.7065 0.9489 0.3985Visualising parallel slopes (from the ANCOVA fit):
lm_ancova <- lm(yield ~ fert + size0, data = ancova)
cols <- c(A = "#1b9e77", B = "#d95f02", C = "#7570b3")
plot(yield ~ size0, data = ancova, col = cols[fert], pch = 19,
main = "ANCOVA: parallel slopes by treatment", xlab = "Baseline size (size0)", ylab = "Yield")
legend("topleft", legend = levels(ancova$fert), col = cols, pch = 19, bty = "n")
slope_hat <- coef(lm_ancova)["size0"]
intercepts <- c(
A = coef(lm_ancova)["(Intercept)"],
B = coef(lm_ancova)["(Intercept)"] + coef(lm_ancova)["fertB"],
C = coef(lm_ancova)["(Intercept)"] + coef(lm_ancova)["fertC"]
)
xg <- seq(min(ancova$size0), max(ancova$size0), length.out = 100)
for (g in names(intercepts)) {
lines(xg, intercepts[g] + slope_hat * xg, col = cols[g], lwd = 2)
}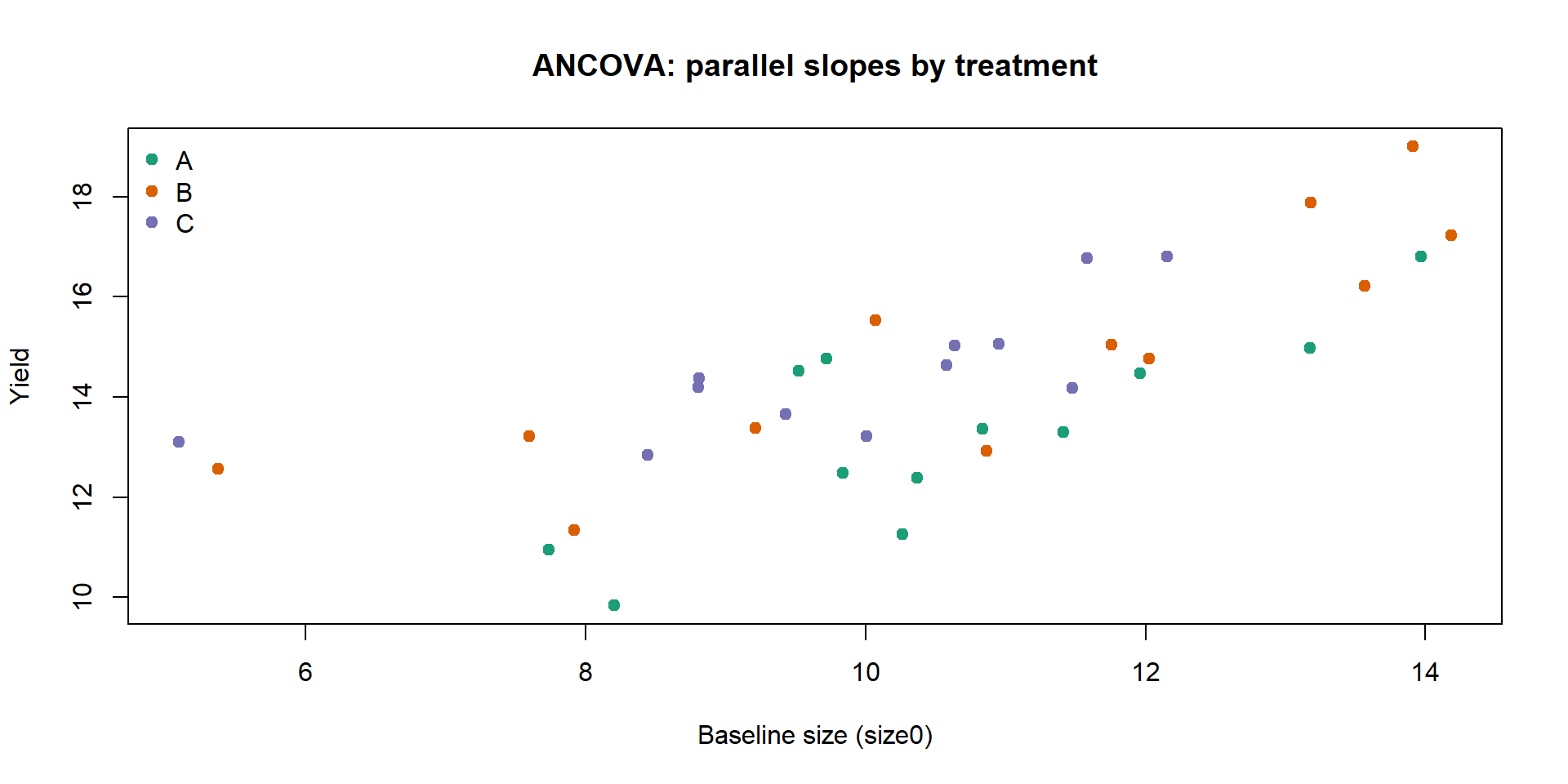
Interpretation:
size0reduces residual variation by explaining baseline differences;ferttests adjusted group shifts.If the
fert:size0interaction is not significant, parallel slopes are a good summary; if significant, the treatment effect depends on the covariate.
Assumptions (in addition to ANOVA’s):
Linearity between the covariate and the response within groups.
Homogeneous slopes across groups (if using the parallel slopes model).
Covariate measured without substantial error and not affected by treatment (pre‑treatment).
MANOVA: multiple responses
Why you need it:
Many experiments record multiple, correlated outcomes (e.g., plant height and leaf area; multiple biochemical markers).
Running separate ANOVAs inflates the familywise error rate and ignores correlation structure.
MANOVA tests for differences in the multivariate mean vector across groups, leveraging correlation to increase power and control Type I error.
What it provides:
A single global test (e.g., Wilks’ Lambda) for whether groups differ on the joint set of responses.
If significant, you can follow up with univariate ANOVAs and multivariate visualisation to understand where differences arise.
set.seed(3)
m <- data.frame(
fert = factor(rep(c("A","B","C"), each = 15))
)
mu <- cbind(A = c(0, 0), B = c(0.8, 0.4), C = c(1.2, 0.9))
Sigma <- matrix(c(1.0, 0.5, 0.5, 1.2), 2, 2) # positive correlation
# If MASS is not available, fall back to independent normals
if (!requireNamespace("MASS", quietly = TRUE)) {
set.seed(3)
y1 <- c(rnorm(15, mu[1, "A"], sqrt(Sigma[1,1])),
rnorm(15, mu[1, "B"], sqrt(Sigma[1,1])),
rnorm(15, mu[1, "C"], sqrt(Sigma[1,1])))
y2 <- c(rnorm(15, mu[2, "A"], sqrt(Sigma[2,2])),
rnorm(15, mu[2, "B"], sqrt(Sigma[2,2])),
rnorm(15, mu[2, "C"], sqrt(Sigma[2,2])))
} else {
set.seed(3)
yA <- MASS::mvrnorm(15, mu = mu[, "A"], Sigma = Sigma)
yB <- MASS::mvrnorm(15, mu = mu[, "B"], Sigma = Sigma)
yC <- MASS::mvrnorm(15, mu = mu[, "C"], Sigma = Sigma)
y1 <- c(yA[,1], yB[,1], yC[,1])
y2 <- c(yA[,2], yB[,2], yC[,2])
}
m$y1 <- y1
m$y2 <- y2
fit_man <- manova(cbind(y1, y2) ~ fert, data = m)
summary(fit_man, test = "Wilks") Df Wilks approx F num Df den Df Pr(>F)
fert 2 0.4634 9.6144 4 82 2e-06 ***
Residuals 42
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Quick visual check of separation:
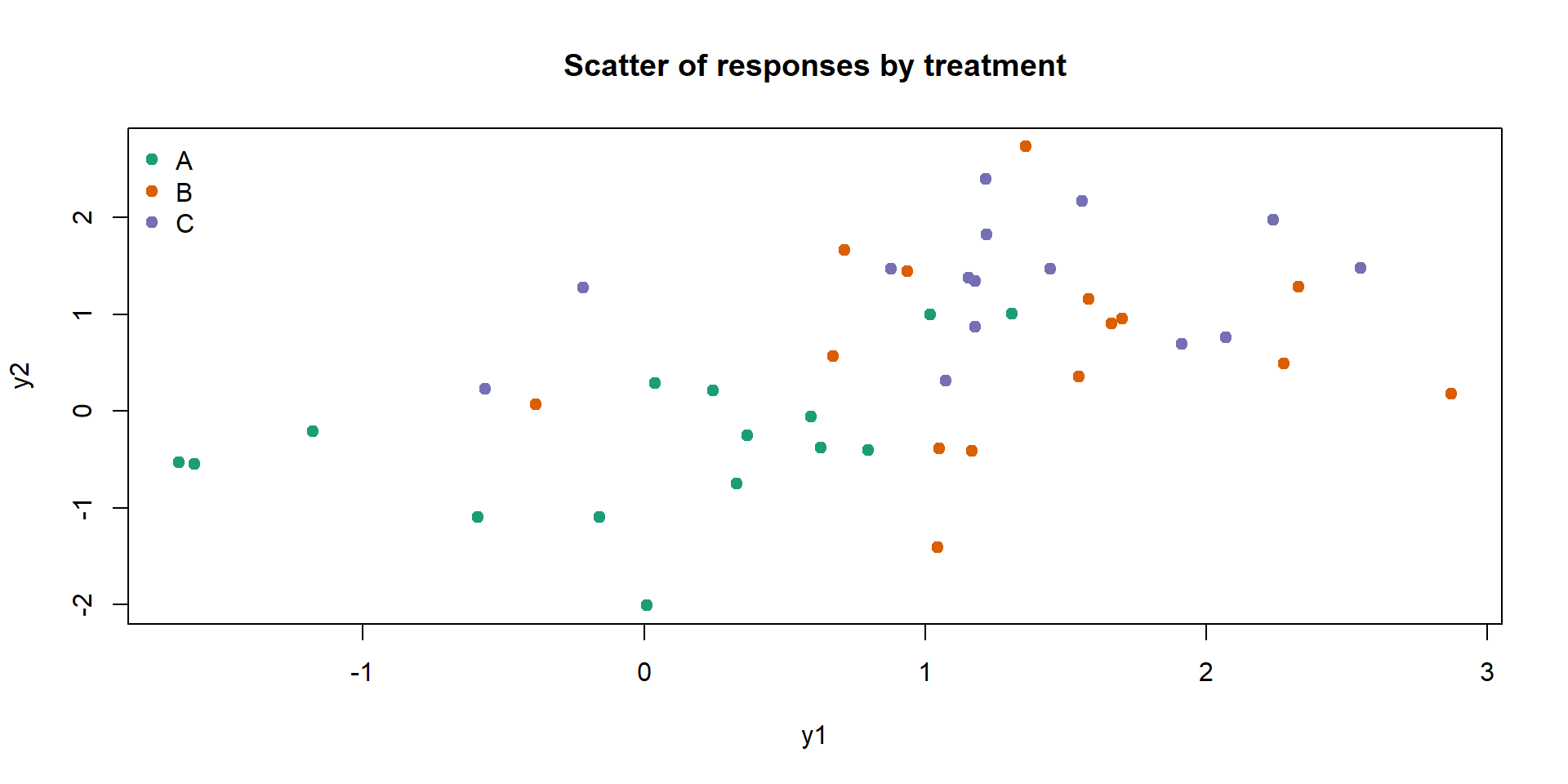
Interpretation and assumptions:
A significant Wilks’ test suggests group centroids differ in the 2D space.
Assumptions: multivariate normality of residuals, equal covariance matrices across groups (Box’s M), and independence. With small samples, be cautious.
Follow‑ups: inspect univariate ANOVAs and confidence ellipses (or discriminant analysis) to interpret which responses drive the separation.
Repeated measures ANOVA (within‑subjects)
Why you need it:
The same subject (or plot) is measured under multiple conditions or times; repeated measures increase efficiency but violate independence.
Repeated measures ANOVA accounts for within‑subject correlation by modelling subjects as a random block and testing the within‑subject factor.
set.seed(4)
subjects <- factor(1:12)
rm_dat <- expand.grid(subject = subjects, treatment = factor(c("A","B","C")))
# Random subject intercepts
subj_eff <- rnorm(length(subjects), 0, 0.8)
rm_dat$response <- with(rm_dat,
5 + (treatment == "B")*0.8 + (treatment == "C")*1.5 +
subj_eff[subject] + rnorm(nrow(rm_dat), 0, 0.5)
)
fit_rm <- aov(response ~ treatment + Error(subject/treatment), data = rm_dat)
summary(fit_rm)
Error: subject
Df Sum Sq Mean Sq F value Pr(>F)
Residuals 11 18.19 1.654
Error: subject:treatment
Df Sum Sq Mean Sq F value Pr(>F)
treatment 2 8.899 4.449 21 7.92e-06 ***
Residuals 22 4.661 0.212
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Simple visualisation of subject trajectories:
cols <- rainbow(length(subjects), alpha = 0.5)
plot(1, type = "n", xlim = c(1, length(levels(rm_dat$treatment))), ylim = range(rm_dat$response),
xaxt = "n", xlab = "Treatment", ylab = "Response", main = "Subject trajectories across treatments")
axis(1, at = 1:length(levels(rm_dat$treatment)), labels = levels(rm_dat$treatment))
for (s in levels(subjects)) {
ds <- subset(rm_dat, subject == s)
ord <- order(ds$treatment)
lines(1:length(ord), ds$response[ord], col = cols[as.integer(s)], lwd = 1.5)
points(1:length(ord), ds$response[ord], col = cols[as.integer(s)], pch = 19)
}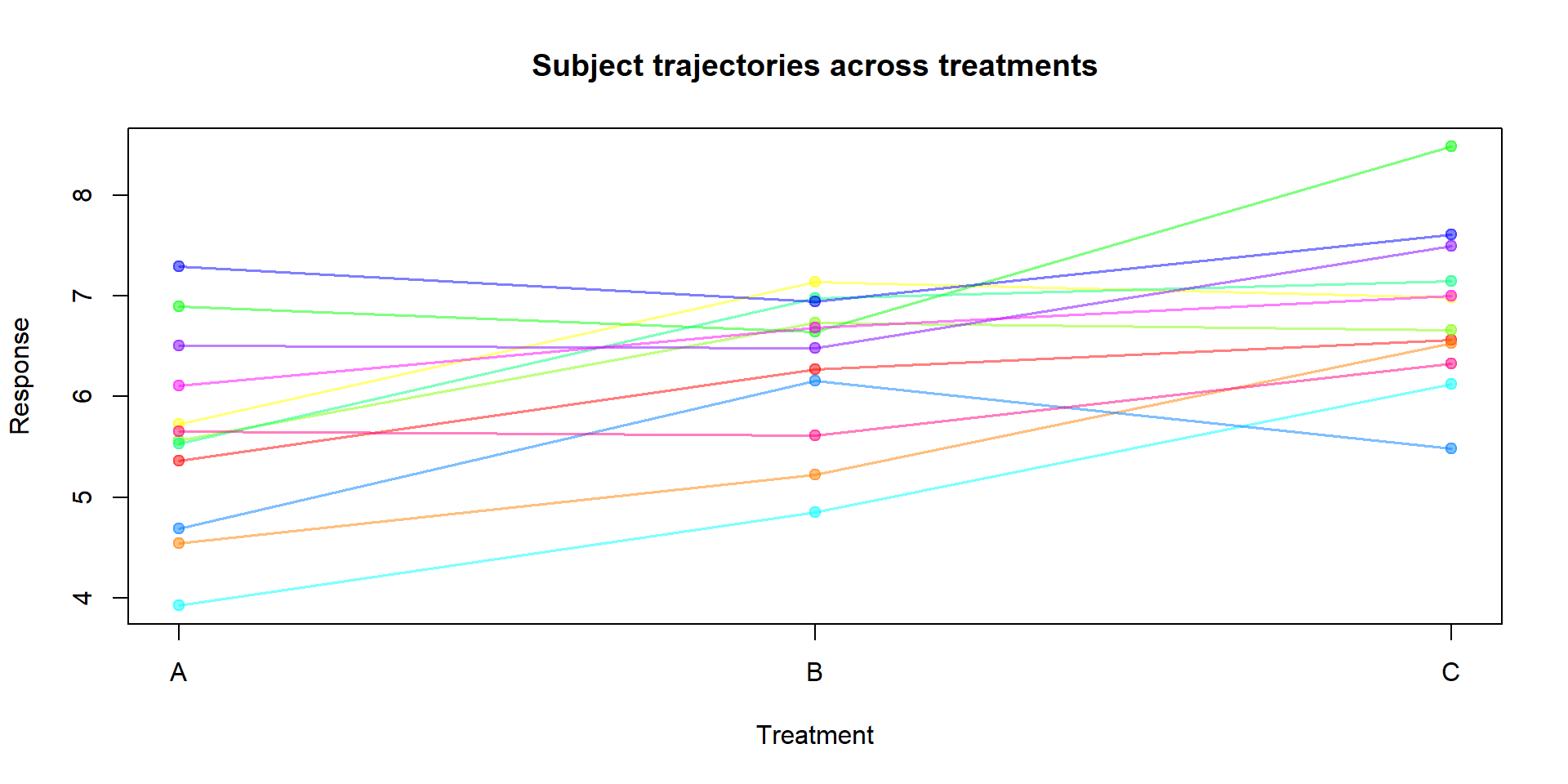
Interpretation and assumptions:
The within‑subject test of
treatmentappears underWithinin theError: subject:treatmentstratum.Sphericity (equal variances of differences between all pairs of levels) is an additional assumption for factors with >2 levels; if violated, corrections (Greenhouse–Geisser, Huynh–Feldt) are used. In more complex designs, consider linear mixed models.
Common pitfalls and tips
Unequal group sizes are fine, but extreme imbalance can reduce power.
Strong outliers can inflate \(SS_W\) or distort group means—investigate them.
With small samples, non‑normal residuals can make p‑values less reliable.
Violated independence (e.g., repeated measures) calls for different models (e.g., mixed effects).
Practice: Try it yourself
Create your own dataset with 4 groups and 6 observations per group. Make two groups similar and two clearly different. Run the full workflow (boxplot → ANOVA → diagnostics → Tukey), and write a two‑sentence report.
Modify the dataset so one group has a much larger variance. What happens to the diagnostics and post‑hoc results?
Build a two‑way dataset with a small block effect. Compare
aov(yield ~ group)vsaov(yield ~ group + block). How does adding blocks change the residual variation and the p‑value forgroup?